|
We have developed a fast geometric algorithm for generating ensemble of protein loop decoys that are free of steric clashes. |
Assumption
|
We assume that an initial loop decoy is given. For a protein with missing loop atoms, standard packages like Modeller can be used to fill in the missing atom coordinates. |
Some Applications
|
|
Some Results
 1(a) |
 1(b) |
|
 1(c) |
 1(d) |
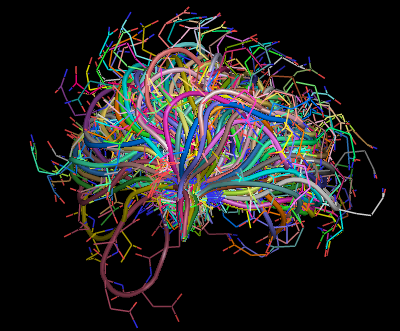 1(e) |
|
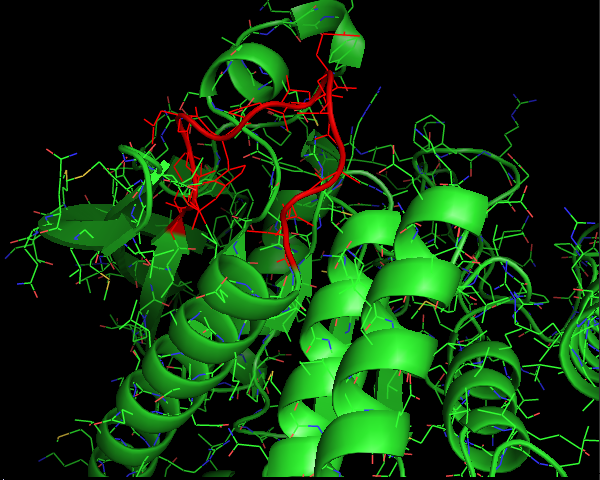
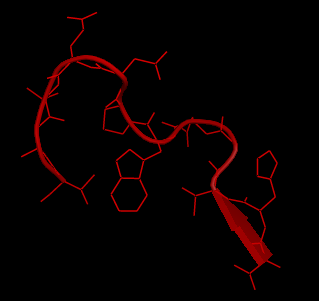
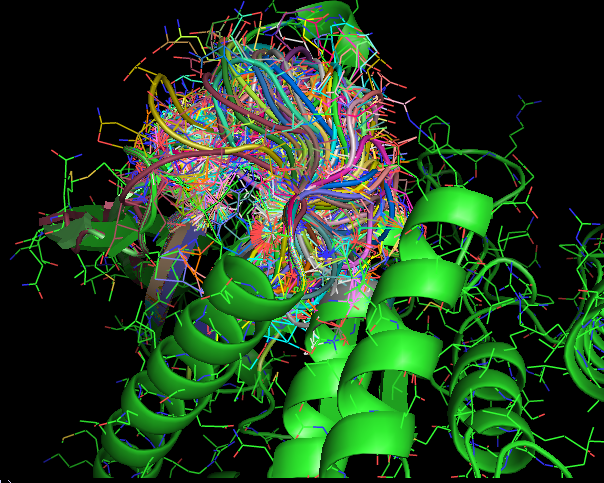
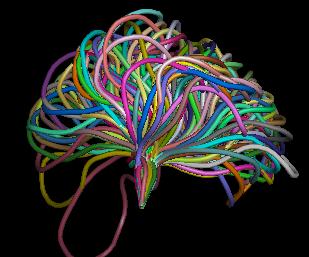
Figure 1(a) and 1(b) show the a 12 residue loop (colored red) on a protein that was chosen. Figures 1(c,d,e) show ensemble of loops that were generated by our technique. All the generated loops are free of steric clashes with respect to itself and the rest of the protein. |
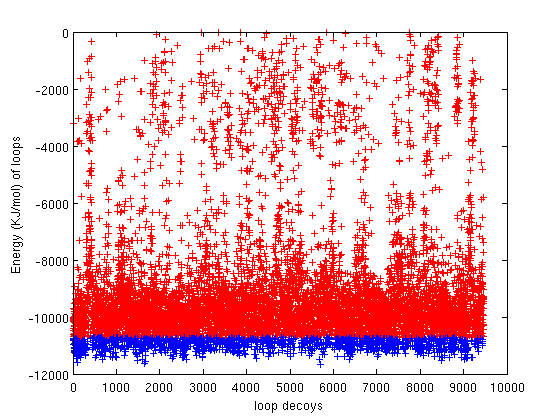 2(a) |
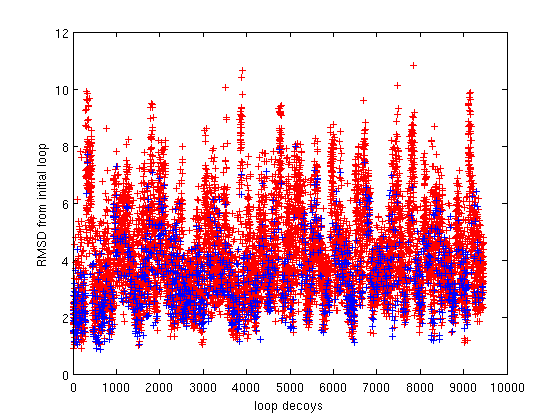 2(b) |
|
Figure 2(a) shows the energy values (y-axis) of the generated loop decoys. Figure 2(b) shows the RMSD of the generated loop decoys with respect to the native loop conoformation. 1255 (blue dots) out of 10000 (red dots) generated decoys were discovered to have better energy than the native conformation. This implies that our technique is very well suited for loop refinement. |
Alogrithm
 |
|
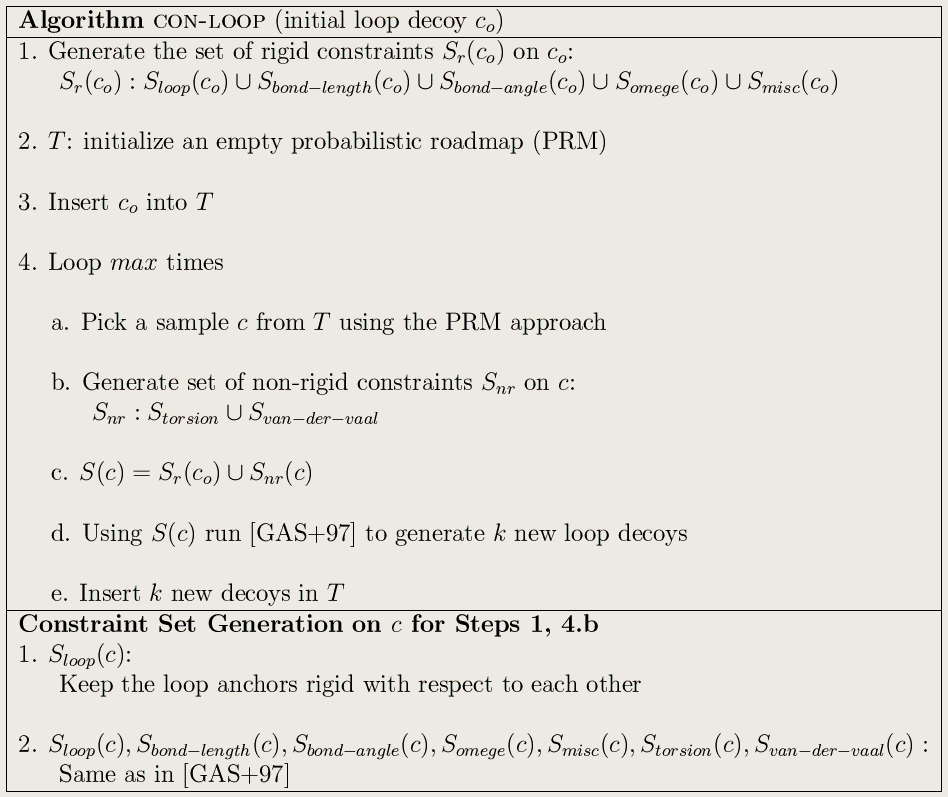
The above figure shows our geometric algorithm for generating loop ensemble.
At each iteration the algorithm randomly perturbs an existing decoy. This will
break the physical constraints (loop constraints, bond length/angle,
steric clashes, etc.) defined the chosen decoy. Then it try's to repair the broken constraints. After the repair it
ends up with a new decoy in the neighborhood of the chosen decoy which satisfies all the constraints.
|
|
|

|
|
|