|
We are studying the stability of RNA folding intermediates using a motion planning approach. We have invented a new geometric sampler for coarse-grained RNA models that can generate low-energy RNA conformations/folding-intermediates which are free from steric clashes and maintain their helical rigidity and tertiary contacts (Figures 2 & 3). This new geometric sampler is used to construct a simplified representation of the RNA conformation space using Probabilistic Roadmaps (PRMs). |
Background
|
Comprised of only four bases, RNA biopolymers adopt surprisingly complex three-dimensional structures and are involved in a wide variety of cellular functions. The observed structural diversity in RNA is a consequence of the diverse hydrogen bonding potential of the four bases and a flexible sugar-phosphate backbone. Most large RNAs are comprised of multiple helices connected by semi-flexible junctions and stabilized by specific tertiary contacts.
|
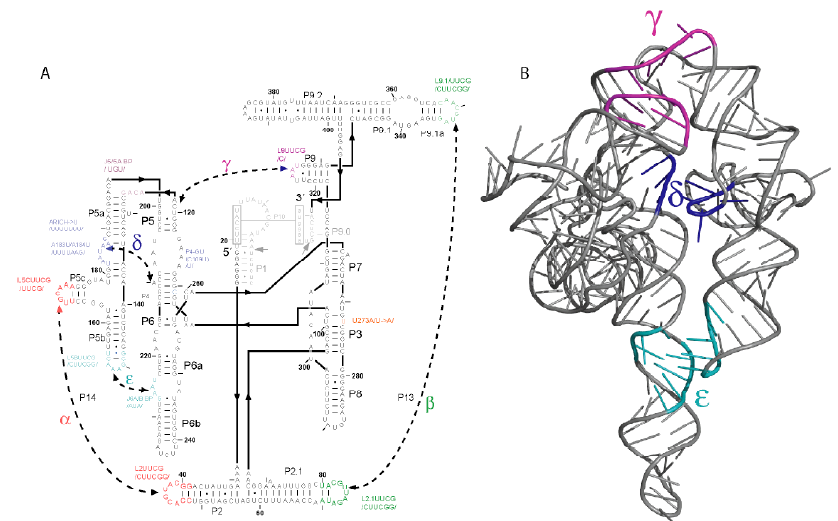 Figure 1 |
|
We define connectivity as the extent to which the formation of one contact affects the stability of another. This term is not to be confounded with macroscopic cooperativity, a thermodynamic quantity generally associated with protein kinetics . In protein biopolymers highly connected networks of tertiary contacts between amino acids, along with a hydrophobic collapse, stabilize the native state. The absence of hydrophobic residues in RNA, and the comparatively small diversity of biopolymer residues (four nucleotides compared to 20 Amino Acids) result in a much sparser tertiary contact map for RNA compared to proteins.
|
Approach and Results
|
We chose as our initial system the P4P6 subdomain of the L-21 T. thermophila group I intron. In Figure 1, this corresponds to nucleotides 104-260. This domain is comprised of 13 helices which pack co-axially, held together by a flexible junction and two tertiary contacts (\delta and \varepsilon). Recent single molecule experiments on this domain have revealed that the cooperativity of the two tertiary contacts is approximately 3 kcal/mol (B. Sattin, personal communication). Therefore, mutating one contact destabilizes the other by approximately 3 kcal/mol. The fundamental question these experiments cannot answer is the origin of these 3 kcal/mol. It is likely that most of the destabilization is due to entropic effects related to the number of conformations available to the system when one tertiary contact is mutated.
|
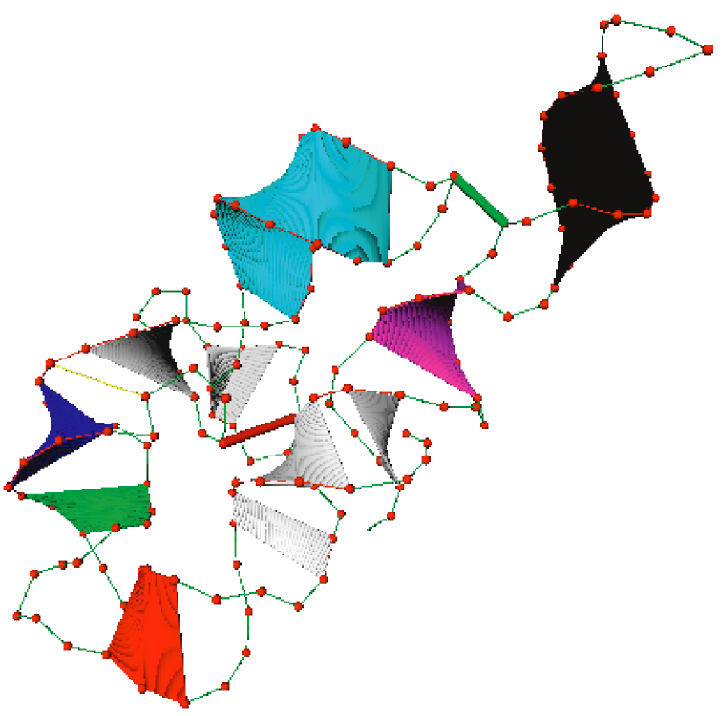 Figure 2: The coarse-grained RNA model. The helical regions (rigid bodies) are shown as sheets and tertiary contacts are shown in red and green thin cylinders. |
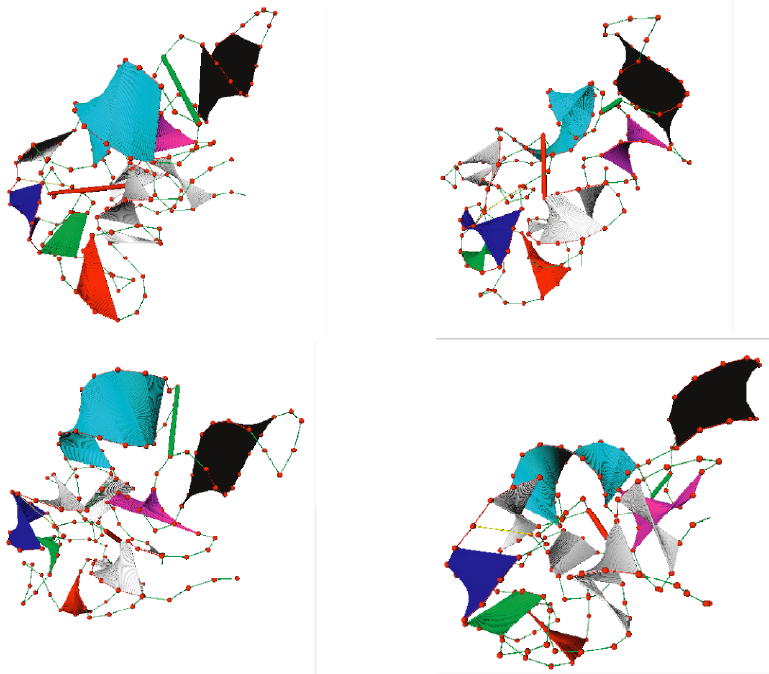 Figure 3: Low energy conformations of the RNA model generated by our new geometric sampler. The desired helical rigidity and tertiary contacts are maintained. |
|
Our approach is to use motion planning to generate alternative conformations of the P4P6 subdomain that are sterically correct given one or both tertiary constraints. We then measure for each helix the total spatial volume given all the conformations generated (in this case, 15,000). Four models were built: In the reference model (Model R, Figure 4), no tertiary constraints were applied and the Volumes of the helices were measured. Two models were built with either the \delta or the \varepsilon constraint (E and D models), and the native model had both constraints (N). We then measured the relative accessible volumes of the helices and report these values in Figure 4. |
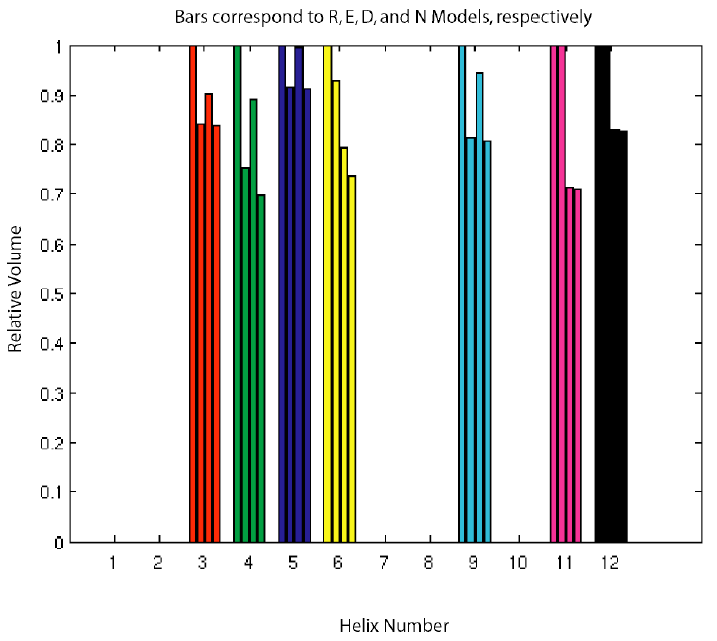 Figure 4 |
|
In Figure 4, the fractional accessible volumes of each helix are shown for the R, E, D, and N models respectively. The values are normalized such that the R (both tertiary contacts missing) model's volumes are always 1.0. As expected, the addition of the tertiary contacts constrains the system and decreases the volumes. The most constrained molecule is in fact the N model. These results therefore demonstrate that we can create a coarse-grained model of RNA and measure the conformational entropy changes due the presence and absence of tertiary contacts. These differences are also spatially specific, as the tertiary contacts affect the relative accessible volumes of the helices differently.
|

|
|
|